From antibodies, nanobodies and peptides to enzymes, membrane proteins, zinc fingers and metalloproteins, Vici.bio helps teams predict structure, optimise binding, edit sequences and run deeper protein calculations in one place.
Predict 3D protein structures, model multimers and support harder cases like protein-ligand, protein-DNA, protein-RNA, metal-bound and cofactor-bound folding with high structural detail.

Evaluate protein-protein docking, compare protein-ligand poses, inspect binding interfaces, map interaction hotspots and support affinity estimation and binding free energy workflows.
Work across humanisation, CDR design, paratope mapping, epitope-aware optimisation, nanobody engineering and peptide binder design for modern therapeutic programs.
Use the editor to mutate residues, scan variants, redesign motifs, tune constructs, adjust linkers and move faster from sequence editing to structural and functional analysis.
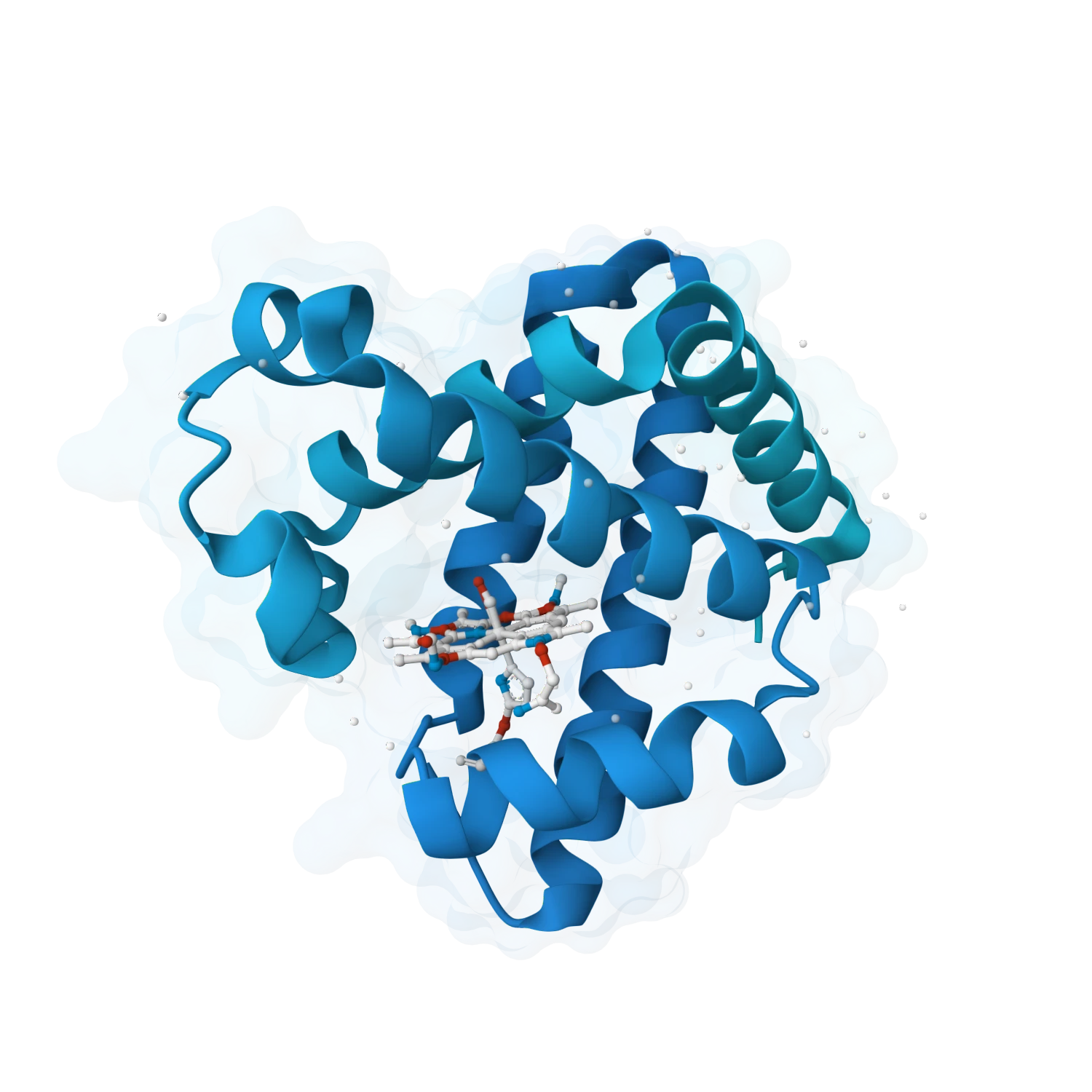
Analyse enzymes, catalytic residues, substrate-binding states, metal coordination, metalloproteins and cofactor-driven systems where chemistry and structure have to be considered together.
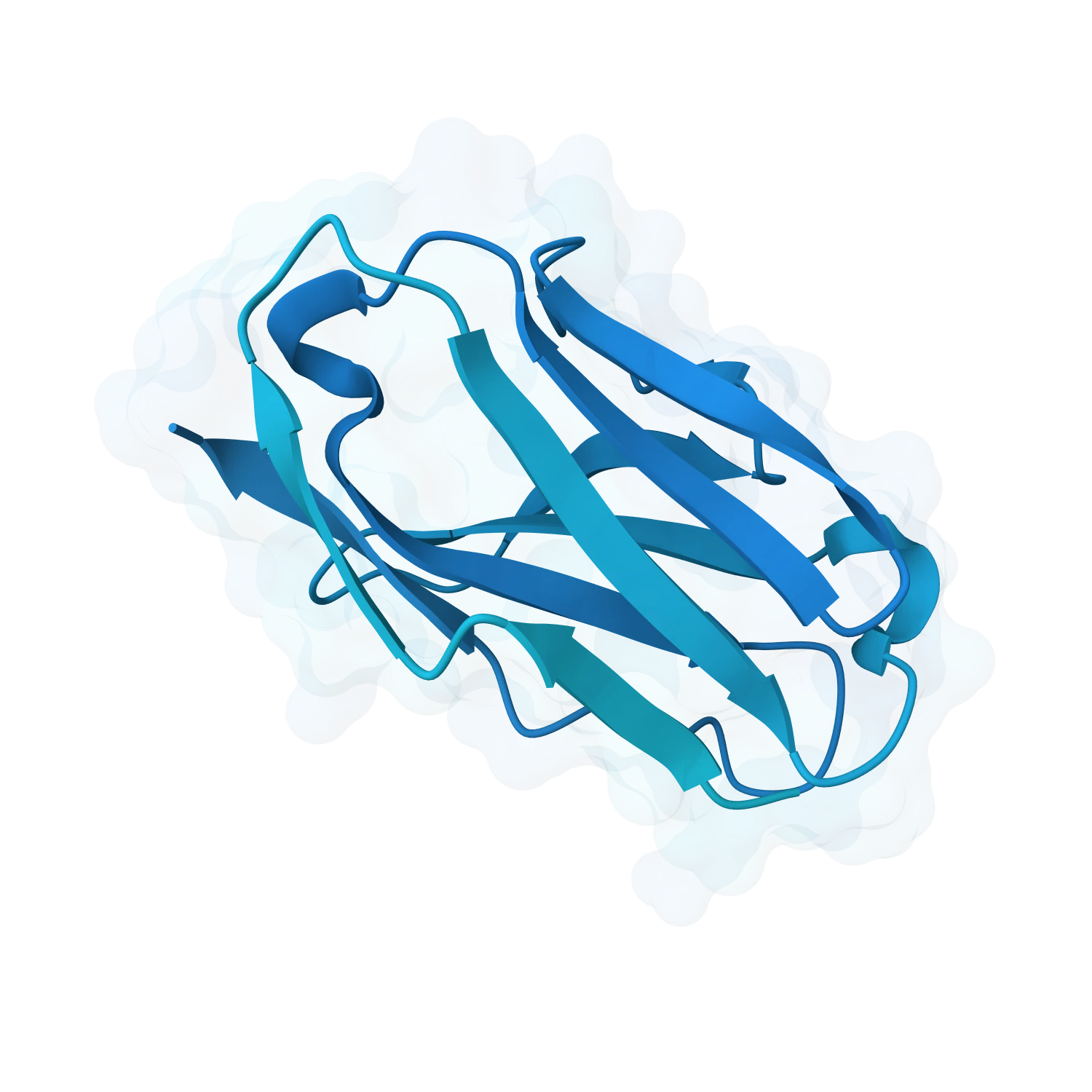
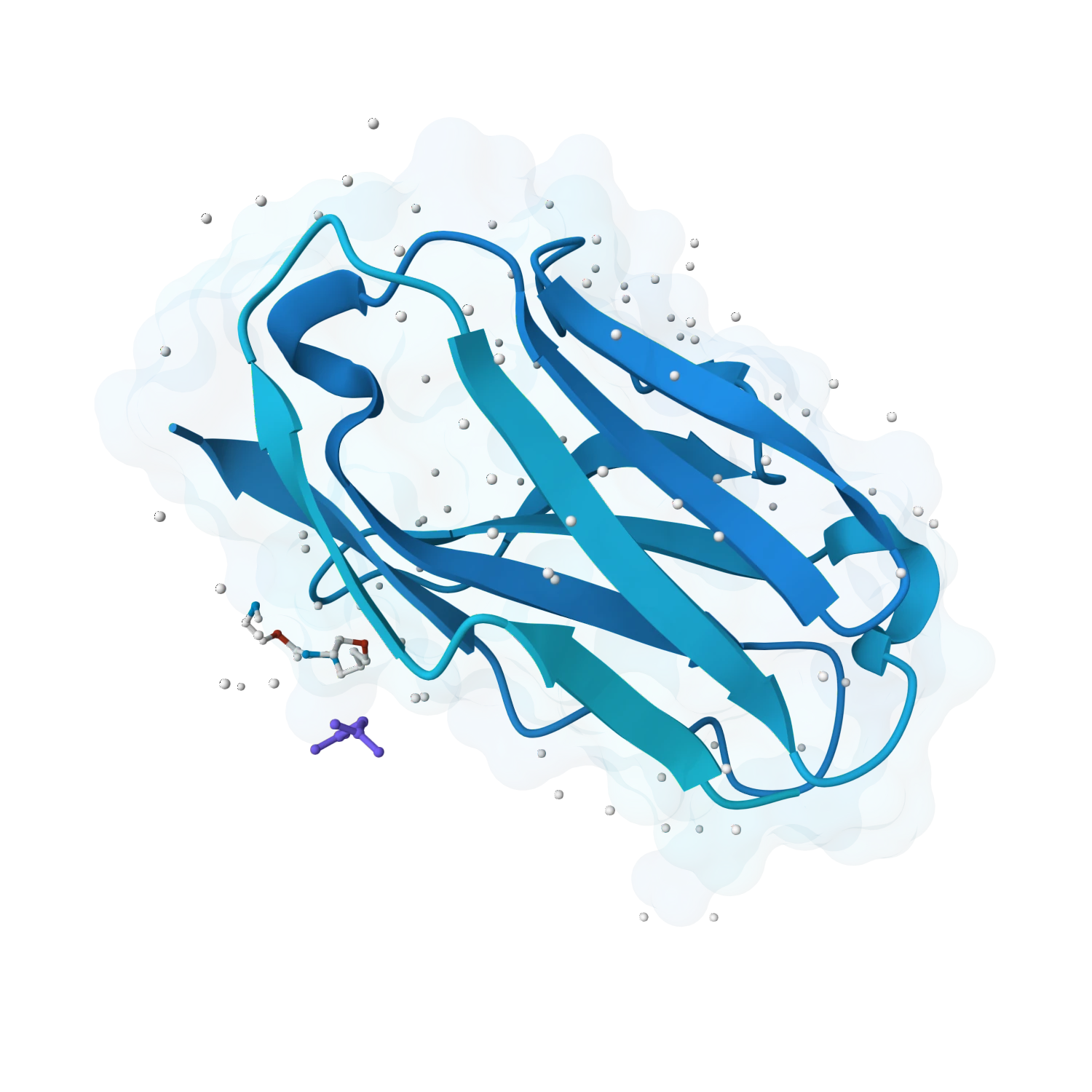
Explore generative protein design for de novo binders, peptides, mini-proteins, repeat proteins and target-specific scaffolds with tools for ranking, optimisation and downstream validation.
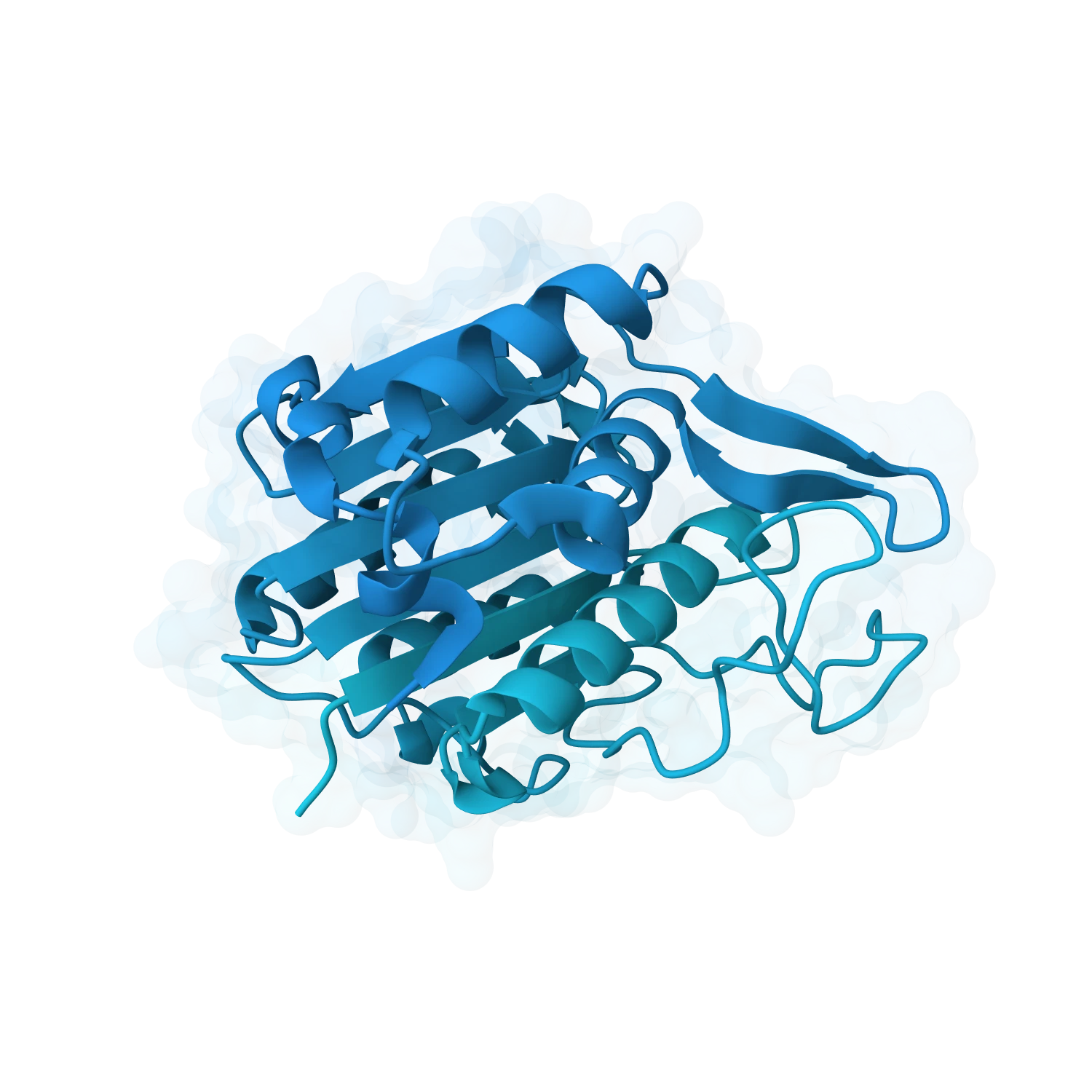
Run molecular dynamics, inspect conformational changes and support harder classes like membrane proteins, intrinsically disordered regions, zinc fingers and larger multimeric assemblies.
From folding and docking to sequence editing, binding energy, molecular dynamics and generative design, build protein workflows with tools designed for modern computational biology.