From small molecules and fragments to glycans, sugars, cyclic dipeptides, macrocycles and other specialised chemical matter, Vici.bio helps teams predict binding, compare poses, estimate affinity and explore deeper ligand-centric calculations.
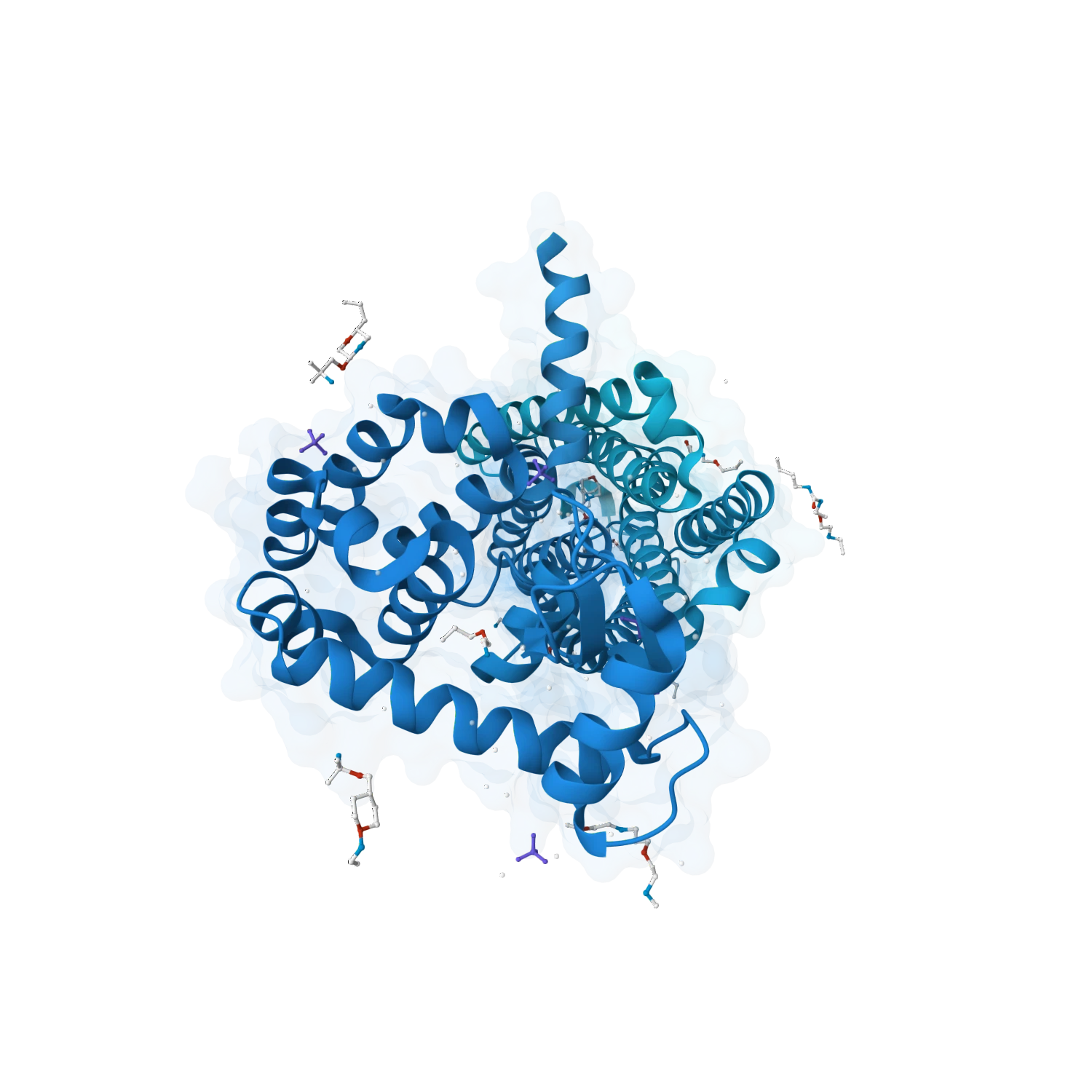
Predict protein-ligand complexes, inspect binding modes and support harder systems involving metals, cofactors, glycans and chemically diverse ligands where context is essential for realistic binding.
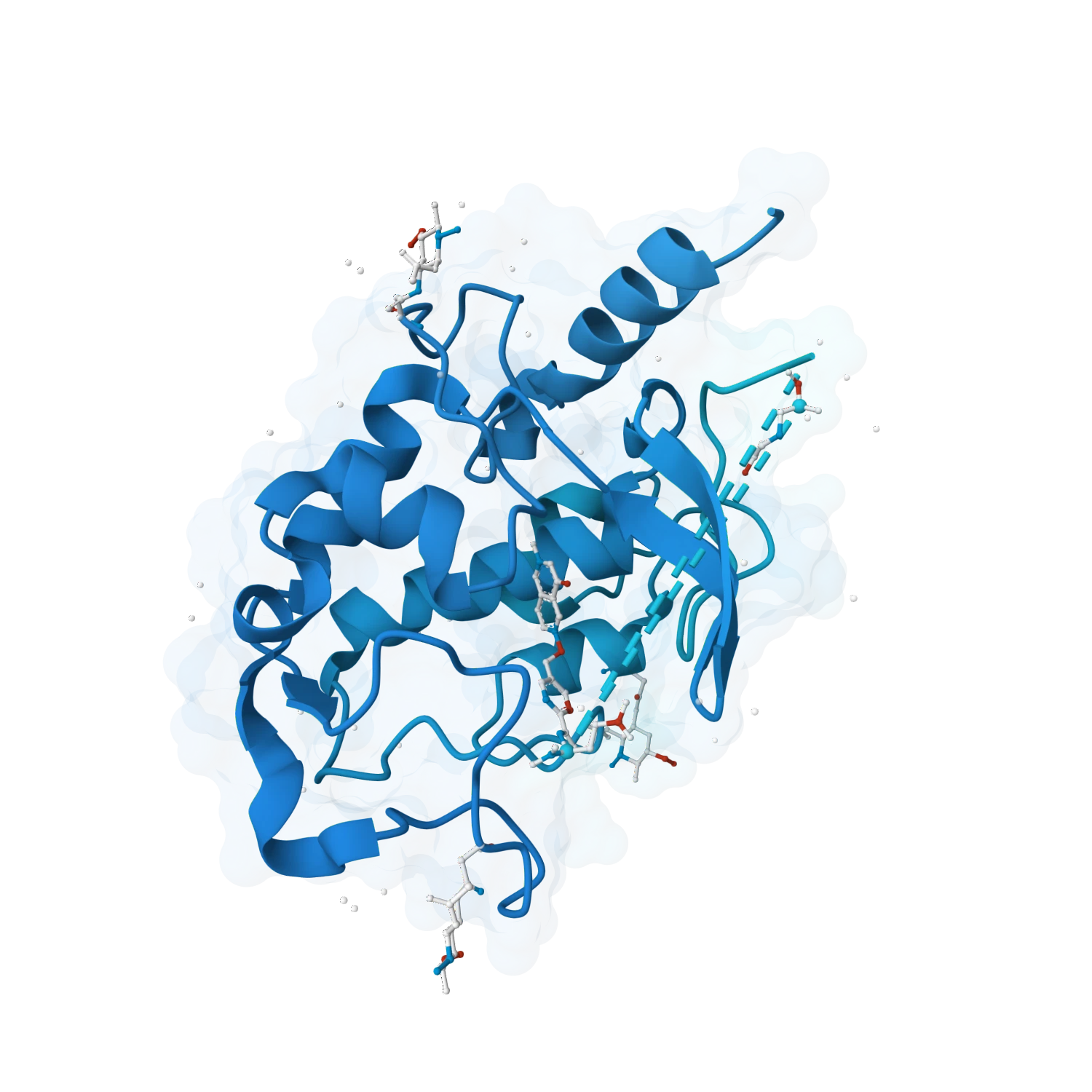
Evaluate ligand docking, compare binding poses, inspect pocket interactions, analyse shape complementarity and rank candidates with cleaner support for screening and lead selection.
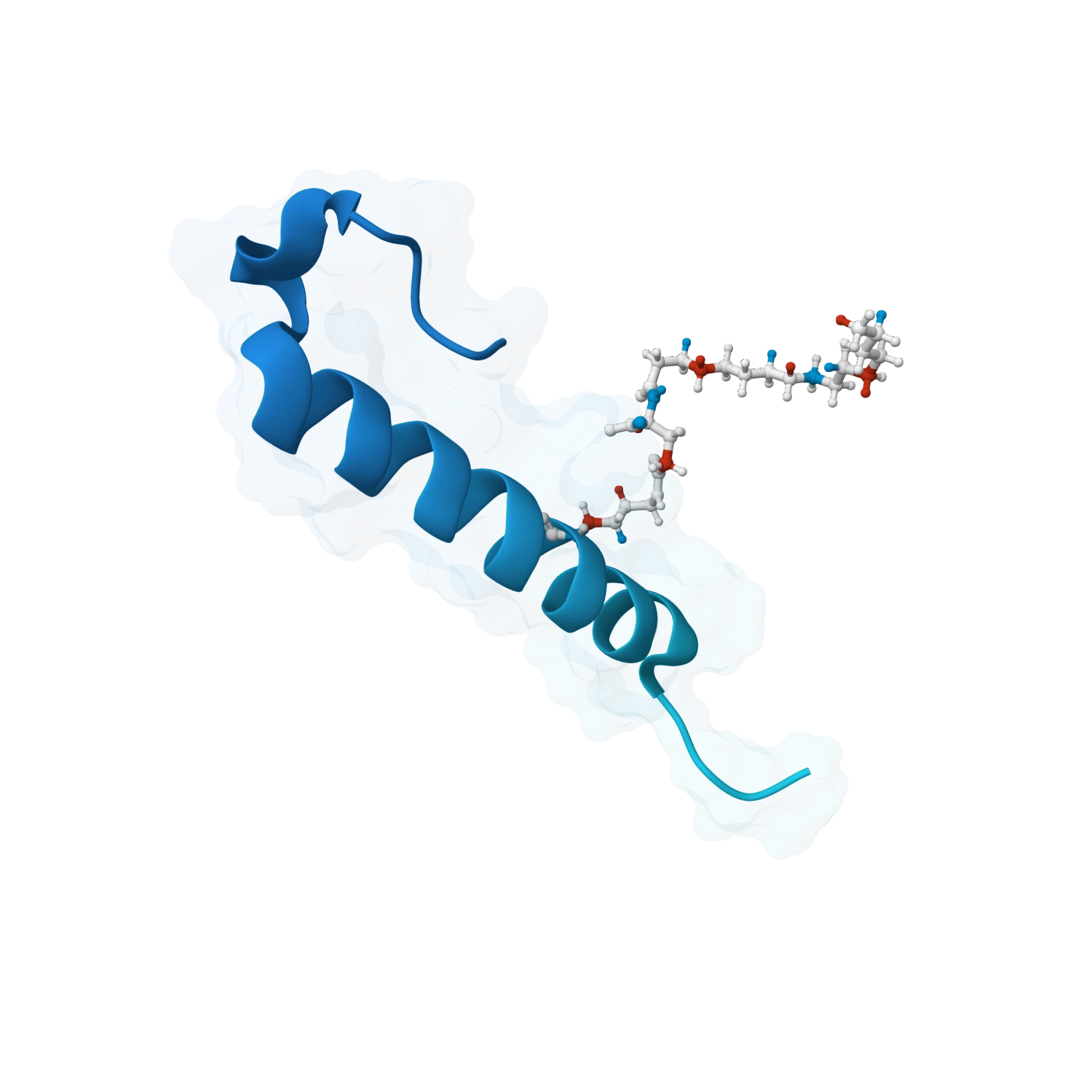
Support affinity prediction, binding energy workflows, interaction scoring and comparative analysis across ligand series so teams can prioritise compounds with stronger evidence, not just better visuals.
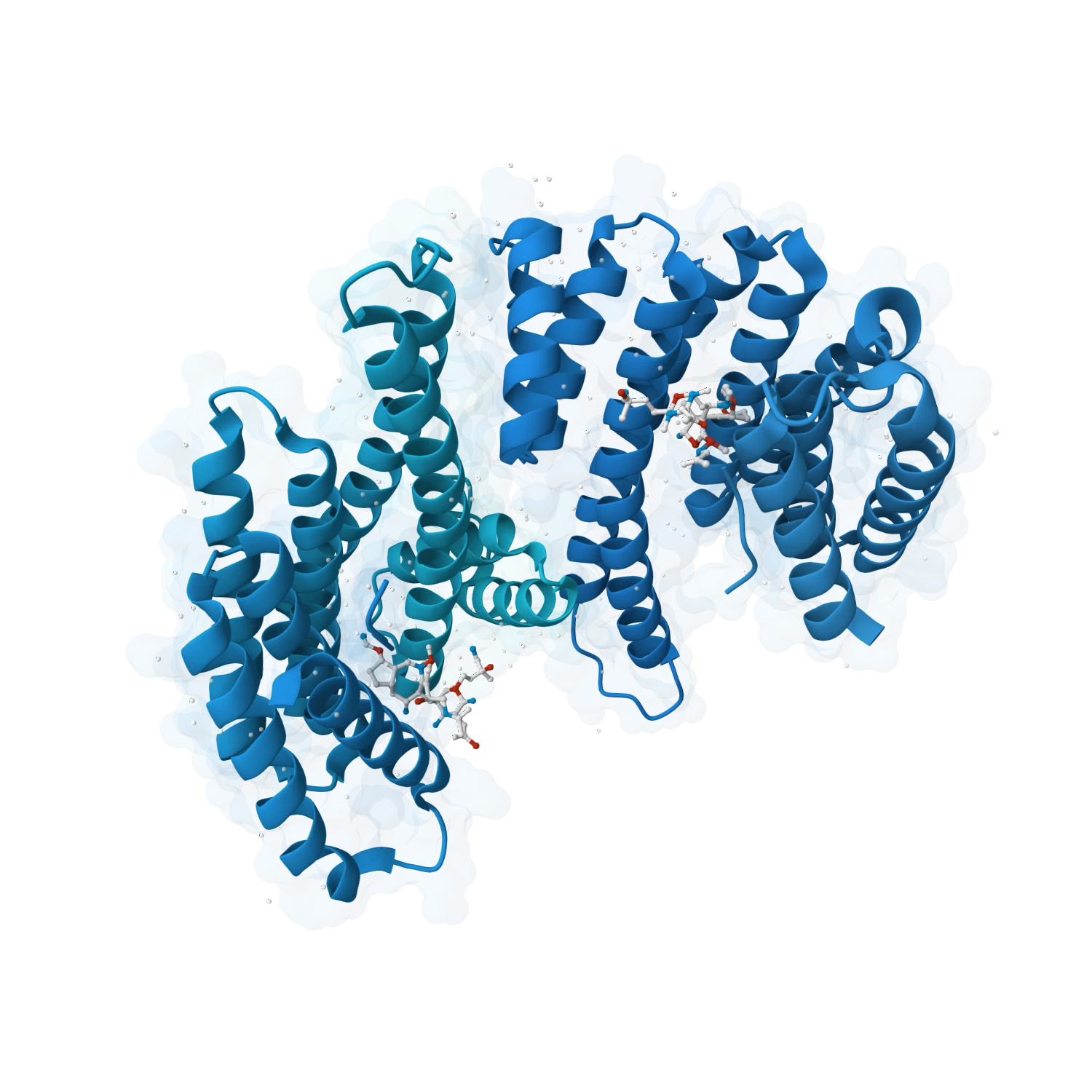
Explore fragment hits, macrocycles, cyclic dipeptides, natural products and more unusual chemotypes that often matter in real discovery programs but are harder to evaluate well.

Support glycans, sugars, carbohydrate ligands, lipid-like molecules and specialised chemical classes where flexibility, stereochemistry and binding context can change the whole interpretation.
Analyse active-site binding, compare substrate-like ligands, inspect catalytic pockets and support workflows where ligand placement, reactivity and local environment all influence compound behaviour.
Explore molecular dynamics, conformational flexibility, induced fit and pocket motion for ligands that cannot be understood from one frozen pose alone, especially in flexible or solvent-exposed systems.
From docking and pose ranking to binding energy, glycan-aware modelling, fragment analysis and molecular dynamics, build ligand workflows designed for modern computational biology.